 Thus, there is a need for robust library production protocols suitable for germline gene inference that satisfy several critical requirements. The current public database for Ig germline genes, the international ImMunoGeneTics information system (IMGT) ( 17), includes alleles from a relatively small number of individuals and thus incompletely covers human global diversity. Two major library production techniques, 5′ Rapid Amplification of cDNA Ends (5′RACE) ( 12, 13) and 5′ multiplex (5′MTPX) PCR ( 14– 16), are used by researchers working with Rep-seq NGS.Īn important first step of Rep-seq analysis, which is required for correct gene assignment and somatic hypermutation (SHM) analysis, is to define the specific germline V alleles present in the subject of interest. 
In order to produce libraries of sufficient length and depth for Rep-seq analysis many groups currently utilize long-read Illumina protocols, such as the 2 × 250 bp HiSeq system or, more commonly, the 2 × 300 bp MiSeq system. This requires a sequence length of at least 400 base pairs (bp), thereby limiting the available sequencing platform options. These libraries are sequenced using NGS protocols that enable the production of amplicons that encompass either partial, in the case of libraries which use framework 1 located primer sequences ( 10, 11), or full-length sequences of the recombined variable (V), diversity (D), and joining (J) gene segments of Ig heavy chains (HC) or VJ sequences of Ig light chains (LC, kappa or lambda). Commonly utilized approaches to immune repertoire sequencing analysis involve the production of isotype-specific libraries of the Ig cDNA. The Adaptive Immune Receptor Repertoire (AIRR) Community are working actively to develop minimum standards and recommendations for repertoire sequencing studies ( 9). The development of NGS-based approaches to Ig repertoire analysis offers new opportunities to investigate B cell responses in health and disease. The library generation protocols presented here enable a robust means of analyzing expressed Ig repertoires, identifying novel alleles and producing individualized germline gene databases from humans. This process additionally uncovered three IGHV, one IGKV, and six IGLV novel alleles in a single individual, which are absent from the IMGT reference database, highlighting the need for further study of Ig genetic variation. Using the optimized protocol, we produced IgM, IgG, IgK, and IgL libraries and analyzed them using the germline inference tool IgDiscover to identify expressed germline V alleles. We provide comprehensive sets of primers targeting IGHV, IGKV, and IGLV genes. We then describe a 5′ multiplex method for library preparation, which yields full length V(D)J sequences suitable for genotype identification and novel gene inference. 
We demonstrate an improved 5′RACE technique that reduces the length constraints of Illumina MiSeq based Rep-seq analysis but allows for the acquisition of sequences upstream of Ig V genes, useful for primer design. Various parameters used in the process were tested in order to demonstrate factors that are critical to obtain high quality libraries. Here, we provide detailed protocols for the production of libraries suitable for human Ig germline gene inference and Ig repertoire studies. All these applications require starting libraries that allow the generation of sequence data with low error rate and optimal representation of the expressed repertoire. Applications include studies of expressed repertoires, gene usage, somatic hypermutation levels, Ig lineage tracing and identification of genetic variation within the Ig loci through inference methods. Next generation sequencing (NGS) of immunoglobulin (Ig) repertoires (Rep-seq) enables examination of the adaptive immune system at an unprecedented level. 3Science for Life Laboratory, Department of Biochemistry and Biophysics, Stockholm University, Stockholm, Sweden.2Division of Rheumatology, Department of Medicine, Center for Molecular Medicine, Karolinska Institutet and Karolinska University Hospital, Stockholm, Sweden.1Department of Microbiology, Tumor and Cell Biology, Karolinska Institutet, Stockholm, Sweden. 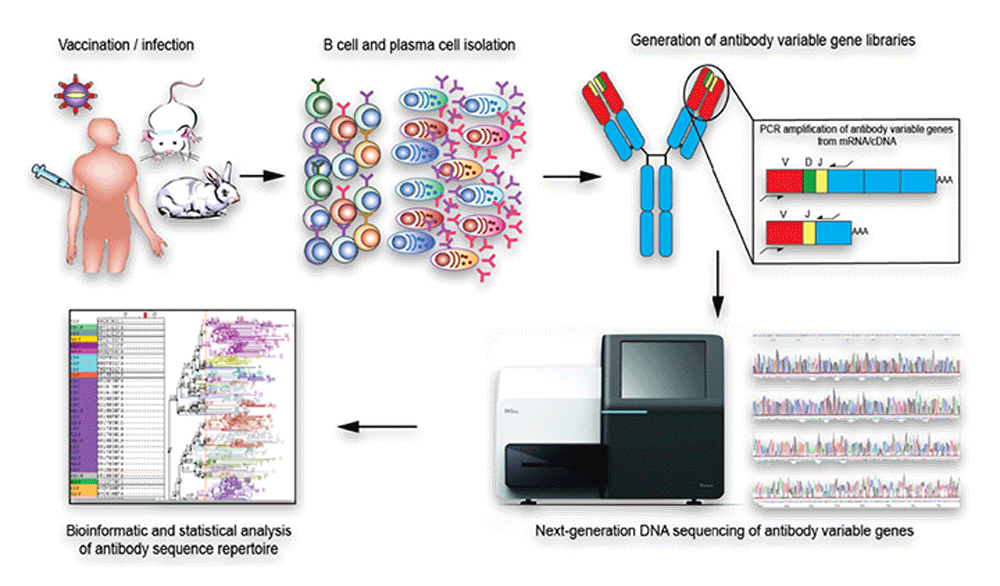
Néstor Vázquez Bernat 1 Martin Corcoran 1 Uta Hardt 1,2 Mateusz Kaduk 1 Ganesh E. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
